Project A3
Predicting evolutionary shifts of cell metabolism
Michael Lässig, U Cologne | web | email
We predict evolutionary changes of bacterial metabolism in response to drug action or environmental challenges. Predictions will be grounded on fitness models that treat external challenge and intrinsic evolutionary response as correlated perturbations of the cell’s metabolic network. The project builds upon previous work on cross-lineage fitness landscapes and involves collaborations with projects A1 and A6.
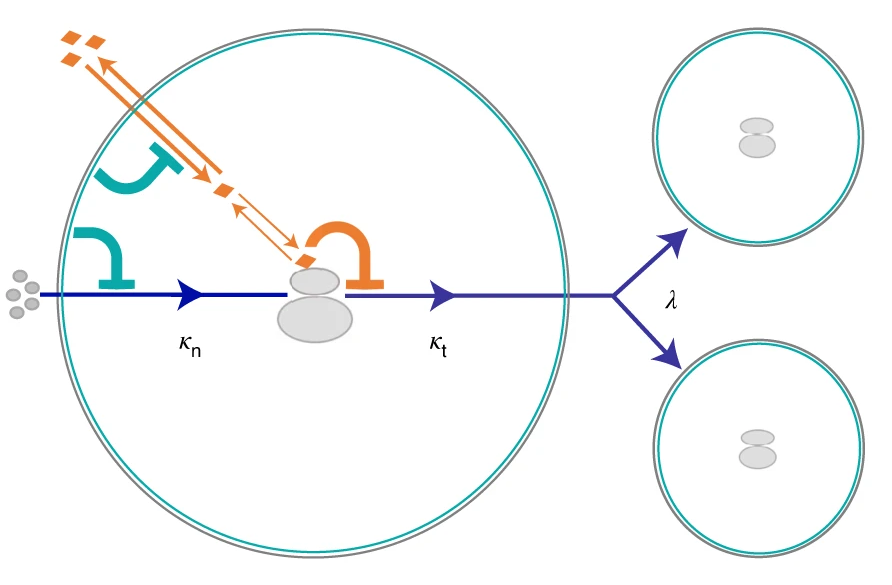
Predictability in Evolution
Collaborative Research Center 1310
Publications
Frazão N., Konrad A., Amicone M., Seixas E., Güleresi D., Lässig M. and Gordo I., Nature Communications, 23. September 2022, https://doi.org/10.1038/s41467-022-33412-8
Metabolic fitness landscapes predict the evolution of antibiotic resistance
Pinheiro, F., Warsi, O., Andersson, D.I., Lässig M., Nat Ecol Evol, 4. March 2021, https://doi.org/10.1038/s41559-021-01397-0
Adaptive evolution of hybrid bacteria by horizontal gene transfer
Power J.J., Pinheiro F., Pompei S., Kovacova V., Yüksel M., Rathmann I., Förster M., Lässig M., Maier B., PNAS 11, 118 (10), 9. March 2021, https://doi.org/10.1073/pnas.2007873118
Horizontal gene transfer overrides mutation in Escherichia coli colonizing the mammalian gut
Frazão N., Sousa A., Lässig M., and Gordo I., PNAS:201906958, 20. August 2019, https://doi.org/10.1073/pnas.1906958116
Survival of the simplest in microbial evolution
Held T., Klemmer D., Lässig M., Nature Communications 10: 2472, 6. June 2019, https://doi.org/10.1038/s41467-019-10413-8
Adaptive Evolution of Gene Expression in Drosophila
Nourmohammad A., Rambeau J., Held T., Kovacova V., Berg J., Lässig M., Cell Reports Volume 20 Issue 6: 1385 - 1395, 8. August 2017, https://doi.org/10.1016/j.celrep.2017.07.033
Lässig M., Mustonen V., Walczak A., Nature Ecol. Evol. 1:0077, 21. February 2017, http://dx.doi.org/10.1038/s41559-017-0077