Project A2
resistance
Joachim Krug, U Cologne | web | email
Arjan de Visser, Wageningen University | web | email
This project has a multi-level research agenda on drug resistance evolution. By a combination of theoretical modeling and laboratory evolution experiments, we aim at making concrete predictions about the evolution of resistance to β-lactam antibiotics. We focus on the relationship between genotype, phenotype, and fitness, including social interactions influencing resistance evolution.
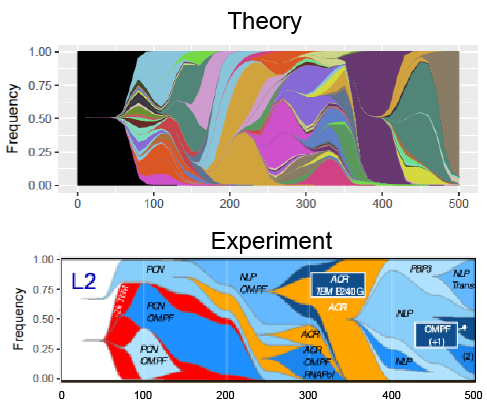
Predictability in Evolution
Collaborative Research Center 1310
Publications
Schenk M.F., Zwart M.P., Hwang S., Ruelens P., Severing E., Krug J., de Visser J.A.G.M., Nat Ecol Evol, 3. March 2022, https://doi.org/10.1038/s41559-022-01669-3
Saebelfeld M., Das S.G., Brink J., Hagenbeek A., Krug J., de Visser J.A.G.M., Frontiers in Microbiology, 12: 698970., 19. Auguts 2021, https://doi.org/10.3389/fmicb.2021.698970
Choice of β-Lactam Resistance Pathway Depends Critically on Initial Antibiotic Concentration
Ruelens P., de Visser J.A.G.M., Antimicrob. Agents Chemother. 65:e00471-21, 16. July 2021, https://doi.org/10.1128/AAC.00471-21
Clonal Interference and Mutation Bias in Small Bacterial Populations in Droplets
Ruelens P., de Visser J.A.G.M., Genes 2021, 12(2), 223, 4. February 2021, https://doi.org/10.3390/genes12020223
The fitness landscapes of translation
Josupeit M., Krug J., Physica A 569, 125768 (2021), 18. January 2021, https://doi.org/10.1016/j.physa.2021.125768
Distribution of the number of fitness maxima in Fisher's geometric model
Park S.C., Hwang S. and Krug J., J. Phys. A: Math. Theor. 53 385601, 21. August 2020, https://doi.org/10.1088/1751-8121/ab9780
Predictable properties of fitness landscapes induced by adaptational tradeoffs
Das S.G., Direito S.O.L., Waclaw B., Allen R.J., Krug J., elife 9, e55155 (2020), 19. May 2020, https://doi.org/10.7554/eLife.55155
Recombination and mutational robustness in neutral fitness landscapes
Klug A., Park S.C, Krug J., PLoS Comput Biol 15(8): e1006884., 15. August 2019, https://doi.org/10.1371/journal.pcbi.1006884
Rare beneficial mutations cannot halt Muller's ratchet in spatial populations
Park S.-C., Klatt P., Krug J., EPL 123: 48001, 3. September 2018, http://stacks.iop.org/0295-5075/123/i=4/a=48001
The utility of fitness landscapes and big data for predicting evolution
de Visser, J. A. G. M., Santiago F. E., Fragata I., Matuszewski S., Heredity 121: 401–405, 20. August 2018, https://doi.org/10.1038/s41437-018-0128-4
Ecology dictates evolution? About the importance of genetic and ecological constraints in adaptation
de Vos M. G. J., Schoustra S. E., de Visser J. A. G. M., EPL 122: 58002, 17. July 2018, http://stacks.iop.org/0295-5075/122/i=5/a=58002
Unraveling the causes of adaptive benefits of synonymous mutations in TEM-1 β-lactamase
Zwart M. P., Schenk M. F., Hwang S., Koopmanschap B., de Lange N., van de Pol L., Nga T. T. T., Szendro I. G., Krug J., de Visser J. A. G. M., Heredity 121: 406–421, 2. July 2018, https://doi.org/10.1038/s41437-018-0104-z
Universality Classes of Interaction Structures for NK Fitness Landscapes
Hwang S., Schmiegelt B., Ferretti L., Krug J., J Stat Phys 172: 226, 13. February 2018, https://doi.org/10.1007/s10955-018-1979-z
Local Fitness Landscapes Predict Yeast Evolutionary Dynamics in Directionally Changing Environments
Gorter F. A., Aarts M. G. M, Zwaan B. J., de Visser J. A. G. M., Genetics 208: 307-322, 1. January 2018, https://doi.org/10.1534/genetics.117.300519
Adaptive benefits from small mutation supplies in an antibiotic resistance enzyme
Salverda M. L. M., Koomen J., Koopmanschap B., Zwart M. P., de Visser J. A. G. M., Proc. Natl. Acad. Sci. USA 114: 12773-12778, 13. November 2017, https://doi.org/10.1073/pnas.1712999114